import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
%matplotlib inlinePandas Fundamentals - I
This notebook was created by Ryan Abernathey (© Copyright 2016-2021) and is part of the An Introduction to Earth and Environmental Data Science textbook.
All content in this book (ie, any files and content in the content/ folder) is licensed under the Creative Commons Attribution-ShareAlike 4.0 International (CC BY-SA 4.0) license.
This notebook demonstrates basic features of the pandas python package.
Pandas is an extension to python, so we first have to import the package before we can run anything else.
Pandas Data Structures: Series
A Series represents a one-dimensional array of data. The main difference between a Series and numpy array is that a Series has an index. The index contains the labels that we use to access the data.
There are many ways to create a Series. We will just show a few.
(Data are from the NASA Planetary Fact Sheet.)
names = ['Mercury', 'Venus', 'Earth']
values = [0.3e24, 4.87e24, 5.97e24]
masses = pd.Series(values, index=names)
massesMercury 3.000000e+23
Venus 4.870000e+24
Earth 5.970000e+24
dtype: float64Series have built in plotting methods.
masses.plot(kind='bar')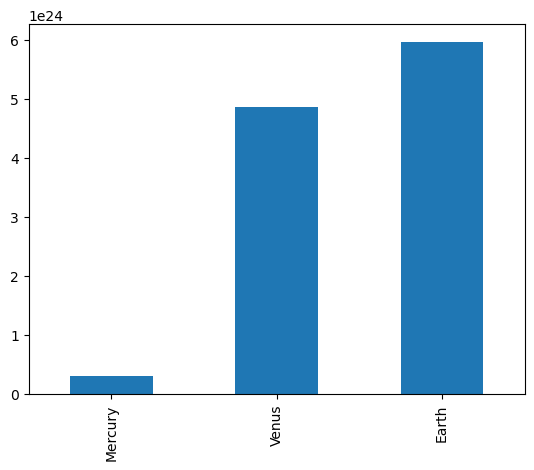
Arithmetic operations and most numpy function can be applied to Series. An important point is that the Series keep their index during such operations.
np.log(masses) / masses**2Mercury 6.006452e-46
Venus 2.396820e-48
Earth 1.600655e-48
dtype: float64We can access the underlying index object if we need to:
masses.indexIndex(['Mercury', 'Venus', 'Earth'], dtype='str')Indexing
We can get values back out using the index via the .loc attribute
masses.loc['Earth']np.float64(5.97e+24)Or by raw position using .iloc
masses.iloc[2]np.float64(5.97e+24)We can pass a list or array to loc to get multiple rows back:
masses.loc[['Venus', 'Earth']]Venus 4.870000e+24
Earth 5.970000e+24
dtype: float64And we can even use slice notation
masses.loc['Mercury':'Earth']Mercury 3.000000e+23
Venus 4.870000e+24
Earth 5.970000e+24
dtype: float64masses.iloc[:2]Mercury 3.000000e+23
Venus 4.870000e+24
dtype: float64If we need to, we can always get the raw data back out as well
masses.values # a numpy arrayarray([3.00e+23, 4.87e+24, 5.97e+24])masses.index # a pandas Index objectIndex(['Mercury', 'Venus', 'Earth'], dtype='str')Pandas Data Structures: DataFrame
There is a lot more to Series, but they are limit to a single “column”. A more useful Pandas data structure is the DataFrame. A DataFrame is basically a bunch of series that share the same index. It’s a lot like a table in a spreadsheet.
Below we create a DataFrame.
# first we create a dictionary
data = {'mass': [0.3e24, 4.87e24, 5.97e24], # kg
'diameter': [4879e3, 12_104e3, 12_756e3], # m
'rotation_period': [1407.6, np.nan, 23.9] # h
}
df = pd.DataFrame(data, index=['Mercury', 'Venus', 'Earth'])
df| mass | diameter | rotation_period | |
|---|---|---|---|
| Mercury | 3.000000e+23 | 4879000.0 | 1407.6 |
| Venus | 4.870000e+24 | 12104000.0 | NaN |
| Earth | 5.970000e+24 | 12756000.0 | 23.9 |
Pandas handles missing data very elegantly, keeping track of it through all calculations.
df.info()<class 'pandas.DataFrame'>
Index: 3 entries, Mercury to Earth
Data columns (total 3 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 mass 3 non-null float64
1 diameter 3 non-null float64
2 rotation_period 2 non-null float64
dtypes: float64(3)
memory usage: 113.0 bytesA wide range of statistical functions are available on both Series and DataFrames.
df.min()mass 3.000000e+23
diameter 4.879000e+06
rotation_period 2.390000e+01
dtype: float64df.mean()mass 3.713333e+24
diameter 9.913000e+06
rotation_period 7.157500e+02
dtype: float64df.std()mass 3.006765e+24
diameter 4.371744e+06
rotation_period 9.784237e+02
dtype: float64df.describe()| mass | diameter | rotation_period | |
|---|---|---|---|
| count | 3.000000e+00 | 3.000000e+00 | 2.000000 |
| mean | 3.713333e+24 | 9.913000e+06 | 715.750000 |
| std | 3.006765e+24 | 4.371744e+06 | 978.423653 |
| min | 3.000000e+23 | 4.879000e+06 | 23.900000 |
| 25% | 2.585000e+24 | 8.491500e+06 | 369.825000 |
| 50% | 4.870000e+24 | 1.210400e+07 | 715.750000 |
| 75% | 5.420000e+24 | 1.243000e+07 | 1061.675000 |
| max | 5.970000e+24 | 1.275600e+07 | 1407.600000 |
We can get a single column as a Series using python’s getitem syntax on the DataFrame object.
df['mass']Mercury 3.000000e+23
Venus 4.870000e+24
Earth 5.970000e+24
Name: mass, dtype: float64…or using attribute syntax.
df.massMercury 3.000000e+23
Venus 4.870000e+24
Earth 5.970000e+24
Name: mass, dtype: float64Indexing works very similar to series
df.loc['Earth']mass 5.970000e+24
diameter 1.275600e+07
rotation_period 2.390000e+01
Name: Earth, dtype: float64df.iloc[2]mass 5.970000e+24
diameter 1.275600e+07
rotation_period 2.390000e+01
Name: Earth, dtype: float64But we can also specify the column we want to access
df.loc['Earth', 'mass']np.float64(5.97e+24)df.iloc[:2, 0]Mercury 3.000000e+23
Venus 4.870000e+24
Name: mass, dtype: float64If we make a calculation using columns from the DataFrame, it will keep the same index:
volume = 4/3 * np.pi * (df.diameter/2)**3
df.mass / volumeMercury 4933.216530
Venus 5244.977070
Earth 5493.285577
dtype: float64Which we can easily add as another column to the DataFrame:
df['density'] = df.mass / volume
df| mass | diameter | rotation_period | density | |
|---|---|---|---|---|
| Mercury | 3.000000e+23 | 4879000.0 | 1407.6 | 4933.216530 |
| Venus | 4.870000e+24 | 12104000.0 | NaN | 5244.977070 |
| Earth | 5.970000e+24 | 12756000.0 | 23.9 | 5493.285577 |
Merging Data
Pandas supports a wide range of methods for merging different datasets. These are described extensively in the documentation. Here we just give a few examples.
temperature = pd.Series([167, 464, 15, -65],
index=['Mercury', 'Venus', 'Earth', 'Mars'],
name='temperature')
temperatureMercury 167
Venus 464
Earth 15
Mars -65
Name: temperature, dtype: int64# returns a new DataFrame
df.join(temperature)| mass | diameter | rotation_period | density | temperature | |
|---|---|---|---|---|---|
| Mercury | 3.000000e+23 | 4879000.0 | 1407.6 | 4933.216530 | 167 |
| Venus | 4.870000e+24 | 12104000.0 | NaN | 5244.977070 | 464 |
| Earth | 5.970000e+24 | 12756000.0 | 23.9 | 5493.285577 | 15 |
# returns a new DataFrame
df.join(temperature, how='right')| mass | diameter | rotation_period | density | temperature | |
|---|---|---|---|---|---|
| Mercury | 3.000000e+23 | 4879000.0 | 1407.6 | 4933.216530 | 167 |
| Venus | 4.870000e+24 | 12104000.0 | NaN | 5244.977070 | 464 |
| Earth | 5.970000e+24 | 12756000.0 | 23.9 | 5493.285577 | 15 |
| Mars | NaN | NaN | NaN | NaN | -65 |
# returns a new DataFrame
everyone = df.reindex(['Mercury', 'Venus', 'Earth', 'Mars'])
everyone| mass | diameter | rotation_period | density | |
|---|---|---|---|---|
| Mercury | 3.000000e+23 | 4879000.0 | 1407.6 | 4933.216530 |
| Venus | 4.870000e+24 | 12104000.0 | NaN | 5244.977070 |
| Earth | 5.970000e+24 | 12756000.0 | 23.9 | 5493.285577 |
| Mars | NaN | NaN | NaN | NaN |
We can also index using a boolean series. This is very useful
adults = df[df.mass > 4e24]
adults| mass | diameter | rotation_period | density | |
|---|---|---|---|---|
| Venus | 4.870000e+24 | 12104000.0 | NaN | 5244.977070 |
| Earth | 5.970000e+24 | 12756000.0 | 23.9 | 5493.285577 |
df['is_big'] = df.mass > 4e24
df| mass | diameter | rotation_period | density | is_big | |
|---|---|---|---|---|---|
| Mercury | 3.000000e+23 | 4879000.0 | 1407.6 | 4933.216530 | False |
| Venus | 4.870000e+24 | 12104000.0 | NaN | 5244.977070 | True |
| Earth | 5.970000e+24 | 12756000.0 | 23.9 | 5493.285577 | True |
Modifying Values
We often want to modify values in a dataframe based on some rule. To modify values, we need to use .loc or .iloc
df.loc['Earth', 'mass'] = 5.98+24
df.loc['Venus', 'diameter'] += 1
df| mass | diameter | rotation_period | density | is_big | |
|---|---|---|---|---|---|
| Mercury | 3.000000e+23 | 4879000.0 | 1407.6 | 4933.216530 | False |
| Venus | 4.870000e+24 | 12104001.0 | NaN | 5244.977070 | True |
| Earth | 2.998000e+01 | 12756000.0 | 23.9 | 5493.285577 | True |
Plotting
DataFrames have all kinds of useful plotting built in.
df.plot(kind='scatter', x='mass', y='diameter', grid=True)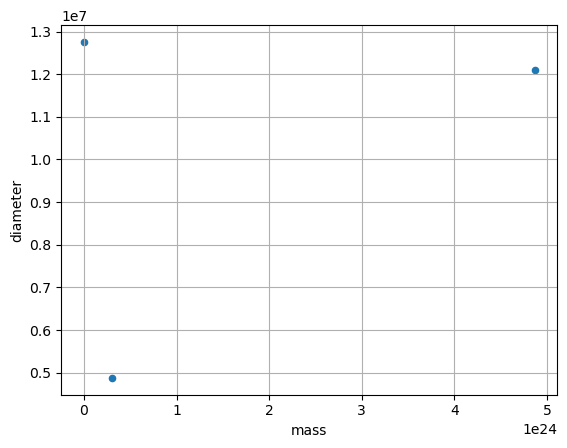
df.plot(kind='bar')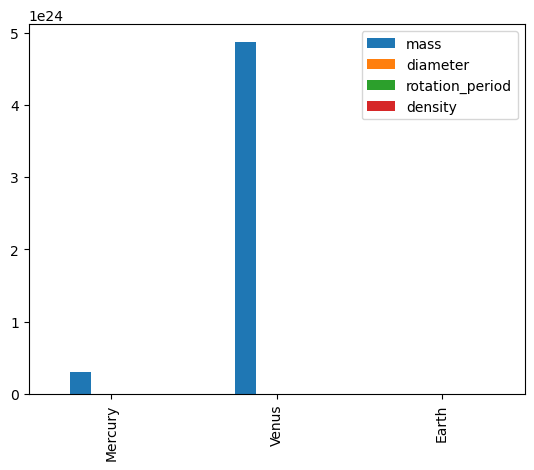
Reading Data Files: Palmer Penguin Data
In this example, we will use data from the Palmer Archipelago Penguins. This dataset has become a popular example dataset for environmental analysis, because it is comparatively simple to understand and has some nice features for analysis. (see e.g. here)
These data were collected from 2007 - 2009 by Dr. Kristen Gorman with the Palmer Station Long Term Ecological Research Program, part of the US Long Term Ecological Research Network. The data are originally from the Environmental Data Initiative (EDI) Data Portal, and are available for use by CC0 license (“No Rights Reserved”) in accordance with the Palmer Station Data Policy.
To read it into pandas, we will use the read_csv function. This function is incredibly complex and powerful. You can use it to extract data from almost any text file. However, you need to understand how to use its various options.
df_penguins = pd.read_csv('../Data/palmer_penguin_data.csv')
df_penguins| Data originally published in: Gorman KB | Williams TD | Fraser WR (2014). Ecological sexual dimorphism and environmental variability within a community of Antarctic penguins (genus Pygoscelis). PLoS ONE 9(3):e90081. https://doi.org/10.1371/journal.pone.0090081 | ||||||
|---|---|---|---|---|---|---|---|---|
| rowid | species | island | bill_length_mm | bill_depth_mm | flipper_length_mm | body_mass_g | sex | year |
| 1 | Adelie | Torgersen | 39.1 | 18.7 | 181 | 3750 | male | 2007 |
| 2 | Adelie | Torgersen | 39.5 | 17.4 | 186 | 3800 | female | 2007 |
| 3 | Adelie | Torgersen | 40.3 | 18 | 195 | 3250 | female | 2007 |
| 4 | Adelie | Torgersen | nan | nan | nan | NaN | NaN | 2007 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 340 | Chinstrap | Dream | 55.8 | 19.8 | 207 | 4000 | male | 2009 |
| 341 | Chinstrap | Dream | 43.5 | 18.1 | 202 | 3400 | female | 2009 |
| 342 | Chinstrap | Dream | 49.6 | 18.2 | 193 | 3775 | male | 2009 |
| 343 | Chinstrap | Dream | 50.8 | 19 | 210 | 4100 | male | 2009 |
| 344 | Chinstrap | Dream | 50.2 | 18.7 | 198 | 3775 | female | 2009 |
345 rows × 3 columns
This does not look right. It looks like that there is an extra line at the top, with some information about the dataset. We also see that there is some data that seems to be missing. If we look directly into the csv, we see that these are indicated with "NA" This is something that we should address directly.
The pd.read_csv() method has many options for ingesting data:
import pandas as pd
df_penguins = pd.read_csv('../Data/palmer_penguin_data.csv',
sep = ',',
na_values='NA',
skiprows= 1,
)
df_penguins
df_penguins.head()| rowid | species | island | bill_length_mm | bill_depth_mm | flipper_length_mm | body_mass_g | sex | year | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | 1 | Adelie | Torgersen | 39.1 | 18.7 | 181.0 | 3750.0 | male | 2007 |
| 1 | 2 | Adelie | Torgersen | 39.5 | 17.4 | 186.0 | 3800.0 | female | 2007 |
| 2 | 3 | Adelie | Torgersen | 40.3 | 18.0 | 195.0 | 3250.0 | female | 2007 |
| 3 | 4 | Adelie | Torgersen | NaN | NaN | NaN | NaN | NaN | 2007 |
| 4 | 5 | Adelie | Torgersen | 36.7 | 19.3 | 193.0 | 3450.0 | female | 2007 |
This looks better
Great. The missing data is now represented by NaN.
What data types did pandas infer?
df_penguins.info()<class 'pandas.DataFrame'>
RangeIndex: 344 entries, 0 to 343
Data columns (total 9 columns):
# Column Non-Null Count Dtype
--- ------ -------------- -----
0 rowid 344 non-null int64
1 species 344 non-null str
2 island 344 non-null str
3 bill_length_mm 342 non-null float64
4 bill_depth_mm 342 non-null float64
5 flipper_length_mm 342 non-null float64
6 body_mass_g 342 non-null float64
7 sex 333 non-null str
8 year 344 non-null int64
dtypes: float64(4), int64(2), str(3)
memory usage: 30.2 KBQuick Statistics
We can calculate basic statistics for columns that are numeric. We can see that this simply skips any columns with text.
df_penguins.describe()| rowid | bill_length_mm | bill_depth_mm | flipper_length_mm | body_mass_g | year | |
|---|---|---|---|---|---|---|
| count | 344.000000 | 342.000000 | 342.000000 | 342.000000 | 342.000000 | 344.000000 |
| mean | 172.500000 | 43.921930 | 17.151170 | 200.915205 | 4201.754386 | 2008.029070 |
| std | 99.448479 | 5.459584 | 1.974793 | 14.061714 | 801.954536 | 0.818356 |
| min | 1.000000 | 32.100000 | 13.100000 | 172.000000 | 2700.000000 | 2007.000000 |
| 25% | 86.750000 | 39.225000 | 15.600000 | 190.000000 | 3550.000000 | 2007.000000 |
| 50% | 172.500000 | 44.450000 | 17.300000 | 197.000000 | 4050.000000 | 2008.000000 |
| 75% | 258.250000 | 48.500000 | 18.700000 | 213.000000 | 4750.000000 | 2009.000000 |
| max | 344.000000 | 59.600000 | 21.500000 | 231.000000 | 6300.000000 | 2009.000000 |
For columns with text, we can get counts on how often each value occurs. Calculating this for numeric columns does not make sense, so we only select the columns that contain text.
text_columns = ['species', 'island', 'sex']
df_penguins[text_columns].value_counts()species island sex
Gentoo Biscoe male 61
female 58
Chinstrap Dream female 34
male 34
Adelie Dream male 28
female 27
Torgersen female 24
male 23
Biscoe female 22
male 22
Name: count, dtype: int64We can now see that there are thee penguin species in the dataset. The Adelie penguin is the only penguin that is found on all three islands (Biscoe, Dream, Torgersen). We also get counts of the sex for each species and island.